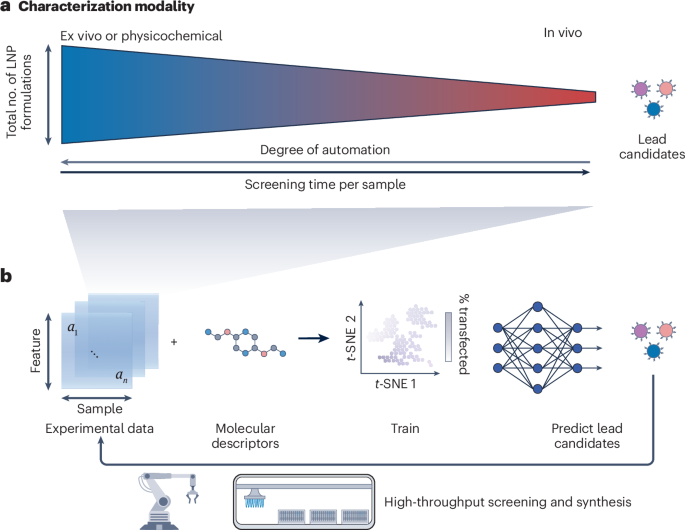
High-throughput platforms for machine learning-guided lipid nanoparticle design
Anderson, D. G., Lynn, D. M. & Langer, R. Semi-automated synthesis and screening of a large library of degradable cationic polymers for gene delivery. Angew. Chem. Int. Ed. 42, 3153–3158 (2003).
Google Scholar
Breunig, M., Lungwitz, U., Liebl, R. & Goepferich, A. Breaking up the correlation between efficacy and toxicity for nonviral gene delivery. Proc. Natl Acad. Sci. USA 104, 14454–14459 (2007).
Google Scholar
Lynn, D. M. & Langer, R. Degradable poly(β-amino esters): synthesis, characterization, and self-assembly with plasmid DNA. J. Am. Chem. Soc. 122, 10761–10768 (2000).
Google Scholar
Siegwart, D. J. et al. Combinatorial synthesis of chemically diverse core-shell nanoparticles for intracellular delivery. Proc. Natl Acad. Sci. USA 108, 12996–13001 (2011).
Google Scholar
Wightman, L. et al. Different behavior of branched and linear polyethylenimine for gene delivery in vitro and in vivo. J. Gene Med. 3, 362–372 (2001).
Google Scholar
Dahlman, J. E. et al. In vivo endothelial siRNA delivery using polymeric nanoparticles with low molecular weight. Nat. Nanotechnol. 9, 648–655 (2014).
Google Scholar
Patra, J. K. et al. Nano based drug delivery systems: recent developments and future prospects. J. Nanobiotechnol. 16, 71 (2018).
Google Scholar
Anselmo, A. C. & Mitragotri, S. Nanoparticles in the clinic: an update. Bioeng. Transl. Med. 4, e10143 (2019).
Google Scholar
Maurer, M. S. et al. Patisiran treatment in patients with transthyretin cardiac amyloidosis. N. Engl. J. Med. 389, 1553–1565 (2023).
Google Scholar
Hassett, K. J. et al. Optimization of lipid nanoparticles for intramuscular administration of mRNA vaccines. Mol. Ther. Nucleic Acids 15, 1–11 (2019).
Google Scholar
Verma, M. et al. The landscape for lipid-nanoparticle-based genomic medicines. Nat. Rev. Drug Discov. 22, 349–350 (2023).
Google Scholar
Dehghani-Ghahnaviyeh, S. et al. Ionizable amino lipids distribution and effects on DSPC/cholesterol membranes: implications for lipid nanoparticle structure. J. Phys. Chem. B 127, 6928–6939 (2023).
Google Scholar
McDonald, S. M. et al. Applied machine learning as a driver for polymeric biomaterials design. Nat. Commun. 14, 4838 (2023).
Google Scholar
Li, B. et al. Accelerating ionizable lipid discovery for mRNA delivery using machine learning and combinatorial chemistry. Nat. Mater. 23, 1002–1008 (2024).
Google Scholar
Han, X. et al. Fast and facile synthesis of amidine-incorporated degradable lipids for versatile mRNA delivery in vivo. Nat. Chem. 16, 1687–1697 (2024).
Google Scholar
Han, X. et al. Optimization of the activity and biodegradability of ionizable lipids for mRNA delivery via directed chemical evolution. Nat. Biomed. Eng. 8, 1412–1424 (2024).
Google Scholar
Xue, L. et al. Combinatorial design of siloxane-incorporated lipid nanoparticles augments intracellular processing for tissue-specific mRNA therapeutic delivery. Nat. Nanotechnol. (2024).
Han, X. et al. Adjuvant lipidoid-substituted lipid nanoparticles augment the immunogenicity of SARS-CoV-2 mRNA vaccines. Nat. Nanotechnol. 18, 1105–1114 (2023).
Google Scholar
Han, X. et al. An ionizable lipid toolbox for RNA delivery. Nat. Commun. 12, 7233 (2021).
Google Scholar
Cheng, Q. et al. Selective organ targeting (SORT) nanoparticles for tissue-specific mRNA delivery and CRISPR–Cas gene editing. Nat. Nanotechnol. 15, 313–320 (2020).
Google Scholar
Dilliard, S. A. & Siegwart, D. J. Passive, active and endogenous organ-targeted lipid and polymer nanoparticles for delivery of genetic drugs. Nat. Rev. Mater. 8, 282–300 (2023).
Google Scholar
Mitchell, M. J. et al. Engineering precision nanoparticles for drug delivery. Nat. Rev. Drug Discov. 20, 101–124 (2021).
Google Scholar
Billingsley, M. M. et al. Orthogonal design of experiments for optimization of lipid nanoparticles for mRNA engineering of CAR T cells. Nano Lett. 22, 533–542 (2022).
Google Scholar
Lokugamage, M. P. et al. Optimization of lipid nanoparticles for the delivery of nebulized therapeutic mRNA to the lungs. Nat. Biomed. Eng. 5, 1059–1068 (2021).
Google Scholar
Hou, X., Zaks, T., Langer, R. & Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Google Scholar
Patel, S. et al. Naturally-occurring cholesterol analogues in lipid nanoparticles induce polymorphic shape and enhance intracellular delivery of mRNA. Nat. Commun. 11, 983 (2020).
Google Scholar
Lokugamage, M. P., Sago, C. D. & Dahlman, J. E. Testing thousands of nanoparticles in vivo using DNA barcodes. Curr. Opin. Biomed. Eng. 7, 1–8 (2018).
Google Scholar
Zheng, L., Bandara, S. R., Tan, Z. & Leal, C. Lipid nanoparticle topology regulates endosomal escape and delivery of RNA to the cytoplasm. Proc. Natl Acad. Sci. USA 120, e2301067120 (2023).
Google Scholar
Koltover, I., Salditt, T., Rädler, J. O. & Safinya, C. R. An inverted hexagonal phase of cationic liposome-DNA complexes related to DNA release and delivery. Science 281, 78–81 (1998).
Google Scholar
Álvarez-Benedicto, E. et al. Optimization of phospholipid chemistry for improved lipid nanoparticle (LNP) delivery of messenger RNA (mRNA). Biomater. Sci. 10, 549–559 (2022).
Google Scholar
Herrera, M., Kim, J., Eygeris, Y., Jozic, A. & Sahay, G. Illuminating endosomal escape of polymorphic lipid nanoparticles that boost mRNA delivery. Biomater. Sci. 9, 4289–4300 (2021).
Google Scholar
Radmand, A. et al. Cationic cholesterol-dependent LNP delivery to lung stem cells, the liver, and heart. Proc. Natl Acad. Sci. USA 121, e2307801120 (2024).
Google Scholar
Mui, B. L. et al. Influence of polyethylene glycol lipid desorption rates on pharmacokinetics and pharmacodynamics of siRNA lipid nanoparticles. Mol. Ther. Nucleic Acids 2, e139 (2013).
Google Scholar
Gindy, M. E. et al. Mechanism of macromolecular structure evolution in self-assembled lipid nanoparticles for siRNA delivery. Langmuir 30, 4613–4622 (2014).
Google Scholar
Kauffman, K. J. et al. Optimization of lipid nanoparticle formulations for mRNA delivery in vivo with fractional factorial and definitive screening designs. Nano Lett. 15, 7300–7306 (2015).
Google Scholar
Akinc, A. et al. A combinatorial library of lipid-like materials for delivery of RNAi therapeutics. Nat. Biotechnol. 26, 561–569 (2008).
Google Scholar
Billingsley, M. M. et al. Ionizable lipid nanoparticle-mediated mRNA delivery for human CAR T cell engineering. Nano Lett. 20, 1578–1589 (2020).
Google Scholar
Whitehead, K. A. et al. Degradable lipid nanoparticles with predictable in vivo siRNA delivery activity. Nat. Commun. 5, 4277 (2014).
Google Scholar
Hajj, K. A. et al. Branched-tail lipid nanoparticles potently deliver mRNA in vivo due to enhanced ionization at endosomal pH. Small 15, 1805097 (2019).
Google Scholar
Yonezawa, S., Koide, H. & Asai, T. Recent advances in siRNA delivery mediated by lipid-based nanoparticles. Adv. Drug Deliv. Rev. 154, 64–78 (2020).
Google Scholar
Li, H. et al. Circular RNA cancer vaccines drive immunity in hard-to-treat malignancies. Theranostics 12, 6422–6436 (2022).
Google Scholar
McKay, P. F. et al. Self-amplifying RNA SARS-CoV-2 lipid nanoparticle vaccine candidate induces high neutralizing antibody titers in mice. Nat. Commun. 11, 3523 (2020).
Google Scholar
Palanki, R. et al. Ionizable lipid nanoparticles for therapeutic base editing of congenital brain disease. ACS Nano 17, 13594–13610 (2023).
Google Scholar
Wei, T. et al. Lung SORT LNPs enable precise homology-directed repair mediated CRISPR/Cas genome correction in cystic fibrosis models. Nat. Commun. 14, 7322 (2023).
Google Scholar
Thatte, A. S. et al. mRNA lipid nanoparticles for ex vivo engineering of immunosuppressive T cells for autoimmunity therapies. Nano Lett. 23, 10179–10188 (2023).
Google Scholar
Billingsley, M. M. et al. In vivo mRNA CAR T cell engineering via targeted ionizable lipid nanoparticles with extrahepatic tropism. Small 20, 2304378 (2024).
Google Scholar
Love, K. T. et al. Lipid-like materials for low-dose, in vivo gene silencing. Proc. Natl Acad. Sci. USA 107, 1864–1869 (2010).
Google Scholar
Forsyth, V., Rzeczycki, P., Taylor, D. & Ariosa, A. Enhancing transfection efficiency of primary immune cells through lipid nanoparticle mediated delivery. J. Immunol. 212, 0857_6526 (2024).
Dilliard, S. A., Cheng, Q. & Siegwart, D. J. On the mechanism of tissue-specific mRNA delivery by selective organ targeting nanoparticles. Proc. Natl Acad. Sci. USA 118, e2109256118 (2021).
Google Scholar
Sun, Y. et al. In vivo editing of lung stem cells for durable gene correction in mice. Science 384, 1196–1202 (2024).
Google Scholar
Qiu, M. et al. Lung-selective mRNA delivery of synthetic lipid nanoparticles for the treatment of pulmonary lymphangioleiomyomatosis. Proc. Natl Acad. Sci. USA 119, e2116271119 (2022).
Google Scholar
LoPresti, S. T., Arral, M. L., Chaudhary, N. & Whitehead, K. A. The replacement of helper lipids with charged alternatives in lipid nanoparticles facilitates targeted mRNA delivery to the spleen and lungs. J. Control. Release 345, 819–831 (2022).
Google Scholar
Omo-Lamai, S. et al. Physicochemical targeting of lipid nanoparticles to the lungs induces clotting: mechanisms and solutions. Adv. Mater. 36, 2312026 (2024).
Google Scholar
Walkey, C. D. et al. Protein corona fingerprinting predicts the cellular interaction of gold and silver nanoparticles. ACS Nano 8, 2439–2455 (2014).
Google Scholar
Walkey, C. D., Olsen, J. B., Guo, H., Emili, A. & Chan, W. C. W. Nanoparticle size and surface chemistry determine serum protein adsorption and macrophage uptake. J. Am. Chem. Soc. 134, 2139–2147 (2012).
Google Scholar
Cai, R. et al. Dynamic intracellular exchange of nanomaterials’ protein corona perturbs proteostasis and remodels cell metabolism. Proc. Natl Acad. Sci. USA 119, e2200363119 (2022).
Google Scholar
Abdelkhaliq, A. et al. Impact of nanoparticle surface functionalization on the protein corona and cellular adhesion, uptake and transport. J. Nanobiotechnol. 16, 70 (2018).
Google Scholar
Baimanov, D. et al. In situ analysis of nanoparticle soft corona and dynamic evolution. Nat. Commun. 13, 5389 (2022).
Google Scholar
Lee, Y. K., Choi, E.-J., Webster, T. J., Kim, S.-H. & Khang, D. Effect of the protein corona on nanoparticles for modulating cytotoxicity and immunotoxicity. Int. J. Nanomed. 10, 97–113 (2015).
Mahmoudi, M., Landry, M. P., Moore, A. & Coreas, R. The protein corona from nanomedicine to environmental science. Nat. Rev. Mater. 8, 422–438 (2023).
Google Scholar
Xu, Y. et al. AGILE platform: a deep learning powered approach to accelerate LNP development for mRNA delivery. Nat. Commun. 15, 6305 (2024).
Google Scholar
Cheng, L. et al. Machine learning elucidates design features of plasmid deoxyribonucleic acid lipid nanoparticles for cell type-preferential transfection. ACS Nano 18, 28735–28747 (2024).
Google Scholar
Dormán, G. The rise, fall and revival of combinatorial chemistry. Nachr. Chem. 70, 70–72 (2022).
Google Scholar
Liu, Y. et al. Ultrastiff metamaterials generated through a multilayer strategy and topology optimization. Nat. Commun. 15, 2984 (2024).
Google Scholar
Ha, C. S. et al. Rapid inverse design of metamaterials based on prescribed mechanical behavior through machine learning. Nat. Commun. 14, 5765 (2023).
Google Scholar
Coley, C. W. et al. A robotic platform for flow synthesis of organic compounds informed by AI planning. Science 365, eaax1566 (2019).
Google Scholar
Hashiba, K. et al. Branching ionizable lipids can enhance the stability, fusogenicity, and functional delivery of mRNA. Small Sci. 3, 2200071 (2023).
Google Scholar
Mukalel, A. J. et al. Oxidized mRNA lipid nanoparticles for in situ chimeric antigen receptor monocyte engineering. Adv. Funct. Mater. 34, 2312038 (2024).
Google Scholar
Riley, R. S. et al. Ionizable lipid nanoparticles for in utero mRNA delivery. Sci. Adv. 7, eaba1028 (2021).
Google Scholar
Chen, J. et al. Combinatorial design of ionizable lipid nanoparticles for muscle-selective mRNA delivery with minimized off-target effects. Proc. Natl Acad. Sci. USA 120, e2309472120 (2023).
Google Scholar
Fenton, O. S. et al. Synthesis and biological evaluation of ionizable lipid materials for the in vivo delivery of messenger RNA to B lymphocytes. Adv. Mater. 29, 1606944 (2017).
Google Scholar
Fenton, O. S. et al. Bioinspired alkenyl amino alcohol ionizable lipid materials for highly potent in vivo mRNA delivery. Adv. Mater. 28, 2939–2943 (2016).
Google Scholar
Hatit, M. Z. C. et al. Nanoparticle stereochemistry-dependent endocytic processing improves in vivo mRNA delivery. Nat. Chem. 15, 508–515 (2023).
Google Scholar
Da Silva Sanchez, A. J. et al. Substituting racemic ionizable lipids with stereopure ionizable lipids can increase mRNA delivery. J. Control. Release 353, 270–277 (2023).
Google Scholar
Li, Z. et al. Enzyme-catalyzed one-step synthesis of ionizable cationic lipids for lipid nanoparticle-based mRNA COVID-19 vaccines. ACS Nano 16, 18936–18950 (2022).
Google Scholar
Jacobsen, E. N. Asymmetric catalysis of epoxide ring-opening reactions. Acc. Chem. Res. 33, 421–431 (2000).
Google Scholar
Li, B. et al. Combinatorial design of nanoparticles for pulmonary mRNA delivery and genome editing. Nat. Biotechnol. 41, 1410–1415 (2023).
Google Scholar
Lee, J. et al. A fully automated platform for photoinitiated RAFT polymerization. Digit. Discov. 2, 219–233 (2023).
Google Scholar
Götz, J. et al. High-throughput synthesis provides data for predicting molecular properties and reaction success. Sci. Adv. 9, eadj2314 (2023).
Google Scholar
Kong, F., Yuan, L., Zheng, Y. F. & Chen, W. Automatic liquid handling for life science: a critical review of the current state of the art. SLAS Technol. 17, 169–185 (2012).
Google Scholar
Burger, B. et al. A mobile robotic chemist. Nature 583, 237–241 (2020).
Google Scholar
Dai, T. et al. Autonomous mobile robots for exploratory synthetic chemistry. Nature 635, 890–897 (2024).
Google Scholar
Steiner, S. et al. Organic synthesis in a modular robotic system driven by a chemical programming language. Science 363, eaav2211 (2019).
Google Scholar
Nambiar, A. M. K. et al. Bayesian optimization of computer-proposed multistep synthetic routes on an automated robotic flow platform. ACS Cent. Sci. 8, 825–836 (2022).
Google Scholar
Shields, B. J. et al. Bayesian reaction optimization as a tool for chemical synthesis. Nature 590, 89–96 (2021).
Google Scholar
Ahneman, D. T., Estrada, J. G., Lin, S., Dreher, S. D. & Doyle, A. G. Predicting reaction performance in C–N cross-coupling using machine learning. Science 360, 186–190 (2018).
Google Scholar
Fromer, J. C. & Coley, C. W. An algorithmic framework for synthetic cost-aware decision making in molecular design. Nat. Comput. Sci. 4, 440–450 (2024).
Google Scholar
Kyranos, J. N., Cai, H., Wei, D. & Goetzinger, W. K. High-throughput high-performance liquid chromatography/mass spectrometry for modern drug discovery. Curr. Opin. Biotechnol. 12, 105–111 (2001).
Google Scholar
Ginsburg-Moraff, C. et al. Integrated and automated high-throughput purification of libraries on microscale. SLAS Technol. 27, 350–360 (2022).
Google Scholar
Su, W.-C. et al. A platform method for simultaneous quantification of lipid and nucleic acid components in lipid nanoparticles. J. Chromatogr. A 1746, 465788 (2025).
Google Scholar
Munjoma, N. et al. High throughput LC-MS platform for large scale screening of bioactive polar lipids in human plasma and serum. J. Proteome Res. 21, 2596–2608 (2022).
Google Scholar
Haas, C. P. et al. Open-source chromatographic data analysis for reaction optimization and screening. ACS Cent. Sci. 9, 307–317 (2023).
Google Scholar
Herck, J. V. et al. Operator-independent high-throughput polymerization screening based on automated inline NMR and online SEC. Digit. Discov. 1, 519–526 (2022).
Google Scholar
Ammini, G. D., Hooker, J. P., Herck, J. V., Kumar, A. & Junkers, T. Comprehensive high-throughput screening of photopolymerization under light intensity variation using inline NMR monitoring. Polym. Chem. 14, 2708–2716 (2023).
Google Scholar
Hu, G. & Qiu, M. Machine learning-assisted structure annotation of natural products based on MS and NMR data. Nat. Prod. Rep. 40, 1735–1753 (2023).
Google Scholar
Kulkarni, J. A. et al. On the formation and morphology of lipid nanoparticles containing ionizable cationic lipids and siRNA. ACS Nano 12, 4787–4795 (2018).
Google Scholar
Maeki, M. et al. Understanding the formation mechanism of lipid nanoparticles in microfluidic devices with chaotic micromixers. PLoS ONE 12, e0187962 (2017).
Google Scholar
Jung, H. N., Lee, S.-Y., Lee, S., Youn, H. & Im, H.-J. Lipid nanoparticles for delivery of RNA therapeutics: current status and the role of in vivo imaging. Theranostics 12, 7509–7531 (2022).
Google Scholar
Strelkova Petersen, D. M., Chaudhary, N., Arral, M. L., Weiss, R. M. & Whitehead, K. A. The mixing method used to formulate lipid nanoparticles affects mRNA delivery efficacy and organ tropism. Eur. J. Pharm. Biopharm. 192, 126–135 (2023).
Google Scholar
Padilla, M. et al. Solution biophysics identifies lipid nanoparticle non-sphericity, polydispersity, and dependence on internal ordering for efficacious mRNA delivery. Preprint at bioRxiv (2025).
Fan, Y. et al. Automated high-throughput preparation and characterization of oligonucleotide-loaded lipid nanoparticles. Int. J. Pharm. 599, 120392 (2021).
Google Scholar
Valente, I., Celasco, E., Marchisio, D. L. & Barresi, A. A. Nanoprecipitation in confined impinging jets mixers: production, characterization and scale-up of pegylated nanospheres and nanocapsules for pharmaceutical use. Chem. Eng. Sci. 77, 217–227 (2012).
Google Scholar
He, Z. et al. Size-controlled lipid nanoparticle production using turbulent mixing to enhance oral DNA delivery. Acta Biomater. 81, 195–207 (2018).
Google Scholar
Shepherd, S. J. et al. Throughput-scalable manufacturing of SARS-CoV-2 mRNA lipid nanoparticle vaccines. Proc. Natl Acad. Sci. USA 120, e2303567120 (2023).
Google Scholar
Shepherd, S. J. et al. Scalable mRNA and siRNA lipid nanoparticle production using a parallelized microfluidic device. Nano Lett. 21, 5671–5680 (2021).
Google Scholar
Hammel, M. et al. Correlating the structure and gene silencing activity of oligonucleotide-loaded lipid nanoparticles using small-angle X-ray scattering. ACS Nano 17, 11454–11465 (2023).
Google Scholar
Valencia, P. M., Farokhzad, O. C., Karnik, R. & Langer, R. Microfluidic technologies for accelerating the clinical translation of nanoparticles. Nat. Nanotechnol. 7, 623–629 (2012).
Google Scholar
Kroth, I., Karimov, M. & Karongo, R. Automated preparation of oligonucleotide-loaded lipid nanoparticles using Andrew+TM pipetting robot for high-throughput in-vitro screening. Waters (2024).
Metzger, L. & Kind, M. On the mixing in confined impinging jet mixers — time scale analysis and scale-up using CFD coarse-graining methods. Chem. Eng. Res. Des. 109, 464–476 (2016).
Google Scholar
Shepherd, S. J., Issadore, D. & Mitchell, M. J. Microfluidic formulation of nanoparticles for biomedical applications. Biomaterials 274, 120826 (2021).
Google Scholar
Sreenivasan, K. R. & Antonia, R. A. The phenomenology of small-scale turbulence. Annu. Rev. Fluid Mech. 29, 435–472 (1997).
Google Scholar
Feng, J., Markwalter, C. E., Tian, C., Armstrong, M. & Prud’homme, R. K. Translational formulation of nanoparticle therapeutics from laboratory discovery to clinical scale. J. Transl. Med. 17, 200 (2019).
Google Scholar
Wilhelm, E., Neumann, C., Duttenhofer, T., Pires, L. & Rapp, B. E. Connecting microfluidic chips using a chemically inert, reversible, multichannel chip-to-world-interface. Lab Chip 13, 4343–4351 (2013).
Google Scholar
Ripoll, M. et al. Optimal self-assembly of lipid nanoparticles (LNP) in a ring micromixer. Sci. Rep. 12, 9483 (2022).
Google Scholar
Kimura, N. et al. Development of a microfluidic-based post-treatment process for size-controlled lipid nanoparticles and application to siRNA delivery. ACS Appl. Mater. Interfaces 12, 34011–34020 (2020).
Google Scholar
Chen, D. et al. Rapid discovery of potent siRNA-containing lipid nanoparticles enabled by controlled microfluidic formulation. J. Am. Chem. Soc. 134, 6948–6951 (2012).
Google Scholar
Belliveau, N. M. et al. Microfluidic synthesis of highly potent limit-size lipid nanoparticles for in vivo delivery of siRNA. Mol. Ther. Nucleic Acids 1, e37 (2012).
Google Scholar
Leung, A. K. K., Tam, Y. Y. C., Chen, S., Hafez, I. M. & Cullis, P. R. Microfluidic mixing: a general method for encapsulating macromolecules in lipid nanoparticle systems. J. Phys. Chem. B 119, 8698–8706 (2015).
Google Scholar
Williams, M. S., Longmuir, K. J. & Yager, P. A practical guide to the staggered herringbone mixer. Lab Chip 8, 1121–1129 (2008).
Google Scholar
Longwell, S. A. & Fordyce, P. M. micrIO: an open-source autosampler and fraction collector for automated microfluidic input–output. Lab Chip 20, 93–106 (2019).
Google Scholar
Hanna, A. R. et al. Automated and parallelized microfluidic generation of large and precisely-defined lipid nanoparticle libraries. Preprint at bioRxiv (2025).
Nag, K. et al. DoE-derived continuous and robust process for manufacturing of pharmaceutical-grade wide-range LNPs for RNA-vaccine/drug delivery. Sci. Rep. 12, 9394 (2022).
Google Scholar
McLeod, E. et al. High-throughput and label-free single nanoparticle sizing based on time-resolved on-chip microscopy. ACS Nano 9, 3265–3273 (2015).
Google Scholar
Graewert, M. A. et al. Quantitative size-resolved characterization of mRNA nanoparticles by in-line coupling of asymmetrical-flow field-flow fractionation with small angle X-ray scattering. Sci. Rep. 13, 15764 (2023).
Google Scholar
Li, S. et al. Payload distribution and capacity of mRNA lipid nanoparticles. Nat. Commun. 13, 5561 (2022).
Google Scholar
Li, S. et al. Single-particle spectroscopic chromatography reveals heterogeneous RNA loading and size correlations in lipid nanoparticles. ACS Nano 18, 15729–15743 (2024).
Google Scholar
Penders, J. et al. Single particle automated Raman trapping analysis. Nat. Commun. 9, 4256 (2018).
Google Scholar
Lowenthal, M. S., Antonishek, A. S. & Phinney, K. W. Quantification of mRNA in lipid nanoparticles using mass spectrometry. Anal. Chem. 96, 1214–1222 (2024).
Google Scholar
Medina, J. et al. Omic-scale high-throughput quantitative LC–MS/MS approach for circulatory lipid phenotyping in clinical research. Anal. Chem. 95, 3168–3179 (2023).
Google Scholar
Kinsey, C. et al. Determination of lipid content and stability in lipid nanoparticles using ultra high-performance liquid chromatography in combination with a corona charged aerosol detector. Electrophoresis 43, 1091–1100 (2022).
Google Scholar
Cui, H. et al. LUMI-lab: a foundation model-driven autonomous platform enabling discovery of new ionizable lipid designs for mRNA delivery. Preprint at bioRxiv (2025).
Maharjan, R., Kim, K. H., Lee, K., Han, H.-K. & Jeong, S. H. Machine learning-driven optimization of mRNA-lipid nanoparticle vaccine quality with XGBoost/Bayesian method and ensemble model approaches. J. Pharm. Anal. 14, 100996 (2024).
Google Scholar
Stach, E. et al. Autonomous experimentation systems for materials development: a community perspective. Matter 4, 2702–2726 (2021).
Google Scholar
Land, O., Seider, W. D. & Lee, D. Convolutional neural network augmented soft-sensor for autonomous microfluidic production of uniform bubbles. Chem. Eng. J. 499, 156494 (2024).
Google Scholar
Szymanski, N. J. et al. An autonomous laboratory for the accelerated synthesis of novel materials. Nature 624, 86–91 (2023).
Google Scholar
Chen, J. et al. Navigating phase diagram complexity to guide robotic inorganic materials synthesis. Nat. Synth. 3, 606–614 (2024).
Google Scholar
Kusne, A. G. et al. On-the-fly closed-loop materials discovery via Bayesian active learning. Nat. Commun. 11, 5966 (2020).
Google Scholar
Jan, E. et al. High-content screening as a universal tool for fingerprinting of cytotoxicity of nanoparticles. ACS Nano 2, 928–938 (2008).
Google Scholar
Frei, A. P. et al. Highly multiplexed simultaneous detection of RNAs and proteins in single cells. Nat. Methods 13, 269–275 (2016).
Google Scholar
Krohn-Grimberghe, M. et al. Nanoparticle-encapsulated siRNAs for gene silencing in the haematopoietic stem-cell niche. Nat. Biomed. Eng. 4, 1076–1089 (2020).
Google Scholar
Connors, J. et al. Lipid nanoparticles (LNP) induce activation and maturation of antigen presenting cells in young and aged individuals. Commun. Biol. 6, 188 (2023).
Google Scholar
Haley, R. M. et al. Lipid nanoparticle delivery of small proteins for potent in vivo RAS inhibition. ACS Appl. Mater. Interfaces 15, 21877–21892 (2023).
Google Scholar
Han, E. L. et al. Predictive high-throughput platform for dual screening of mRNA lipid nanoparticle blood-brain barrier transfection and crossing. Nano Lett. 24, 1477–1486 (2024).
Google Scholar
Liu, R., Jiang, W., Walkey, C. D., Chan, W. C. W. & Cohen, Y. Prediction of nanoparticles-cell association based on corona proteins and physicochemical properties. Nanoscale 7, 9664–9675 (2015).
Google Scholar
Alabi, C. A. et al. Multiparametric approach for the evaluation of lipid nanoparticles for siRNA delivery. Proc. Natl Acad. Sci. USA 110, 12881–12886 (2013).
Google Scholar
Roosa, C. A. et al. Conjugation of IL-33 to microporous annealed particle scaffolds enhances type 2-like immune responses in vitro and in vivo. Adv. Healthc. Mater. 13, 2400249 (2024).
Google Scholar
Sahay, G. et al. Efficiency of siRNA delivery by lipid nanoparticles is limited by endocytic recycling. Nat. Biotechnol. 31, 653–658 (2013).
Google Scholar
Rui, Y. et al. High-throughput and high-content bioassay enables tuning of polyester nanoparticles for cellular uptake, endosomal escape, and systemic in vivo delivery of mRNA. Sci. Adv. 8, eabk2855 (2022).
Google Scholar
Ponsoda, X., Jover, R., Castell, J. V. & Gómez-Lechón, M. J. Measurement of intracellular LDH activity in 96-well cultures: a rapid and automated assay for cytotoxicity studies. J. Tissue Cult. Methods 13, 21–24 (1991).
Google Scholar
Marin, D. et al. Safety, efficacy and determinants of response of allogeneic CD19-specific CAR-NK cells in CD19+ B cell tumors: a phase 1/2 trial. Nat. Med. 30, 772–784 (2024).
Google Scholar
Dahlman, J. E. et al. Barcoded nanoparticles for high throughput in vivo discovery of targeted therapeutics. Proc. Natl Acad. Sci. USA 114, 2060–2065 (2017).
Google Scholar
Kim, J., Vaughan, H. J., Zamboni, C. G., Sunshine, J. C. & Green, J. J. High-throughput evaluation of polymeric nanoparticles for tissue-targeted gene expression using barcoded plasmid DNA. J. Control. Release 337, 105–116 (2021).
Google Scholar
Guimaraes, P. P. G. et al. Ionizable lipid nanoparticles encapsulating barcoded mRNA for accelerated in vivo delivery screening. J. Control. Release 316, 404–417 (2019).
Google Scholar
Rhym, L. H., Manan, R. S., Koller, A., Stephanie, G. & Anderson, D. G. Peptide-encoding mRNA barcodes for the high-throughput in vivo screening of libraries of lipid nanoparticles for mRNA delivery. Nat. Biomed. Eng. 7, 901–910 (2023).
Google Scholar
Vaidya, K. et al. Pooled nanoparticle screening using a chemical barcoding approach. Angew. Chem. Int. Ed. 137, e202420052 (2025).
Google Scholar
Boehnke, N. et al. Massively parallel pooled screening reveals genomic determinants of nanoparticle delivery. Science 377, eabm5551 (2022).
Google Scholar
El‐Mayta, R. et al. A nanoparticle platform for accelerated in vivo oral delivery screening of nucleic acids. Adv. Ther. 4, 2000111 (2021).
Google Scholar
Hatit, M. Z. C. et al. Species-dependent in vivo mRNA delivery and cellular responses to nanoparticles. Nat. Nanotechnol. 17, 310–318 (2022).
Google Scholar
Hamilton, A. G. et al. High-throughput in vivo screening identifies differential influences on mRNA lipid nanoparticle immune cell delivery by administration route. ACS Nano 18, 16151–16165 (2024).
Google Scholar
Dobrowolski, C. et al. Nanoparticle single-cell multiomic readouts reveal that cell heterogeneity influences lipid nanoparticle-mediated messenger RNA delivery. Nat. Nanotechnol. 17, 871–879 (2022).
Google Scholar
Zhang, Q. et al. Predictable control of RNA lifetime using engineered degradation-tuning RNAs. Nat. Chem. Biol. 17, 828–836 (2021).
Google Scholar
Bruchez, M., Moronne, M., Gin, P., Weiss, S. & Alivisatos, A. P. Semiconductor nanocrystals as fluorescent biological labels. Science 281, 2013–2016 (1998).
Google Scholar
Paunovska, K. et al. A direct comparison of in vitro and in vivo nucleic acid delivery mediated by hundreds of nanoparticles reveals a weak correlation. Nano Lett. 18, 2148–2157 (2018).
Google Scholar
Sago, C. D. et al. High-throughput in vivo screen of functional mRNA delivery identifies nanoparticles for endothelial cell gene editing. Proc. Natl Acad. Sci. USA 115, E9944–E9952 (2018).
Google Scholar
Buschmann, T. & Bystrykh, L. V. Levenshtein error-correcting barcodes for multiplexed DNA sequencing. BMC Bioinform. 14, 272 (2013).
Google Scholar
Witten, J. et al. Artificial intelligence-guided design of lipid nanoparticles for pulmonary gene therapy. Nat. Biotechnol. (2024).
Cervantes, J., Garcia-Lamont, F., Rodríguez-Mazahua, L. & Lopez, A. A comprehensive survey on support vector machine classification: applications, challenges and trends. Neurocomputing 408, 189–215 (2020).
Google Scholar
Yamankurt, G. et al. Exploration of the nanomedicine-design space with high-throughput screening and machine learning. Nat. Biomed. Eng. 3, 318–327 (2019).
Google Scholar
Kumar, R. et al. Efficient polymer-mediated delivery of gene-editing ribonucleoprotein payloads through combinatorial design, parallelized experimentation, and machine learning. ACS Nano 14, 17626–17639 (2020).
Google Scholar
Pattipeiluhu, R. et al. Anionic lipid nanoparticles preferentially deliver mRNA to the hepatic reticuloendothelial system. Adv. Mater. 34, 2201095 (2022).
Google Scholar
Mandl, H. K. et al. Optimizing biodegradable nanoparticle size for tissue-specific delivery. J. Control. Release 314, 92–101 (2019).
Google Scholar
Popova, M., Isayev, O. & Tropsha, A. Deep reinforcement learning for de novo drug design. Sci. Adv. 4, eaap7885 (2018).
Google Scholar
Tropsha, A., Isayev, O., Varnek, A., Schneider, G. & Cherkasov, A. Integrating QSAR modelling and deep learning in drug discovery: the emergence of deep QSAR. Nat. Rev. Drug Discov. 23, 141–155 (2024).
Google Scholar
Walsh, D. J. et al. Community Resource for Innovation in Polymer Technology (CRIPT): a scalable polymer material data structure. ACS Cent. Sci. 9, 330–338 (2023).
Google Scholar
Kuenneth, C. & Ramprasad, R. polyBERT: a chemical language model to enable fully machine-driven ultrafast polymer informatics. Nat. Commun. 14, 4099 (2023).
Google Scholar
Meyer, T. A., Ramirez, C., Tamasi, M. J. & Gormley, A. J. A user’s guide to machine learning for polymeric biomaterials. ACS Polym. Au 3, 141–157 (2023).
Google Scholar
Xue, K. et al. Biomaterials by design: harnessing data for future development. Mater. Today Bio 12, 100165 (2021).
Google Scholar
Haley, R. M. et al. Lipid nanoparticles for in vivo lung delivery of CRISPR-Cas9 ribonucleoproteins allow gene editing of clinical targets. ACS Nano 19, 13790–13804 (2025).
Google Scholar
Tang, C. et al. mRNA-laden lipid-nanoparticle-enabled in situ CAR-macrophage engineering for the eradication of multidrug-resistant bacteria in a sepsis mouse model. ACS Nano 18, 2261–2278 (2024).
Google Scholar
link






